Rational 3-D Processing Step-By-Step
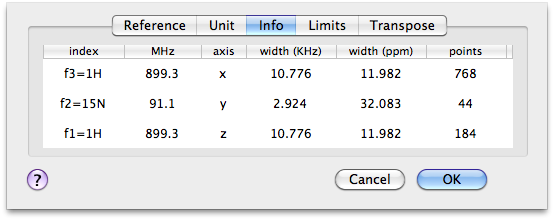
To share my little personal experience, here is how I have processed a 3-D spectrum. Some of the concepts and rules can be applied to other 3-D spectra, others are specific to the example I have been working with. In this example the spectrometer is by Bruker and the pulse sequence is “noesyhsqcfpf3gpsi3d”. Don't panic, it's a NOESY-HSQC experiment. To read the number of points before processing, and orient myself, I use the command: Format > Axes and Scales.
The quality of the spectrum improves if the number of points is at least doubled by zero-filling. For the first FT, I should therefore set the size to at least 1536. The first power of 2 coming after this value is 2048. The other FTs are applied eliminating the imaginary points (because this is a phase-sensitive experiment). The double of the halved sizes are the present sizes: 44 and 184. Rounded to the next powers of 2 they become 64 and 256.
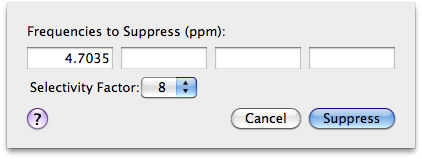
To remove the water signal (or most of it), I choose Process > Suppress. The default frequency is already the middle frequency. All I have to do is to set a lower Selectivity Factor (8 instead of 16): there is no important signal at the center of the spectrum and I want to remove the water as much as I can.
Like most of today's Bruker spectra, this one contains an enigmatic prologue at the left of the FID. We use to call it the digital filter... It doesn't hurt too much, but can be taken into account when choosing a weighting function. In my case, I want to apply a cosine bell, i.e., a sine bell shifted, in theory, by 90°. To make the top of the bell coincide with the beginning of the true FID, a smaller shift is actually required. I have written this short script for the task:
-- CosineBell.lua - calculates the shift corresponding to a cosine bell
FID = getf()
shift = 180 - 90 * FID.size / (FID.size + FID.zero) -- all calculations on this line
print( "The cosine bell corresponds to a shift of ", shift, " degrees" ) -- visual feedback
weight( nil, nil, shift )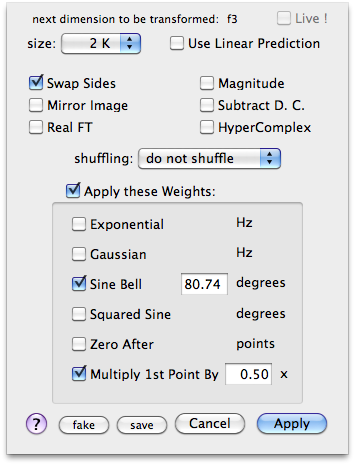
The pulse sequence and the epxeriment itself have been set up in such a way that the FID starts exactly at the end of the pulse sequence. Zero delay = no first-order phase correction is required.
When you aren't going to use first-order phase correction, like in this case, you can use a well-known and simple trick to correct the baseline: multiply the first point of the FID by 0.5. This is how this experiment is supposed to be processed. The principle is that the discrete FT adds a constant bias to the baseline. The trick simply removes the minimal amount of baseline distortion that is inherently present in any FT-NMR spectrum. If we apply the first-order phase correction, the trick cannot work.
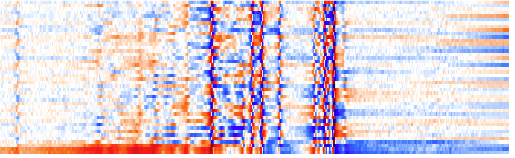
I can correct the phase directly on the whole matrix (but paying attention to the bottom row eclusively). When there is too much color the phase is wrong; the whiter, the better. No need to be accurate now, I will refine the phase later on. I will never use the first-order correction on this spectrum, for the reason explained before.
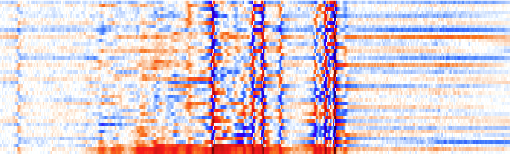
After phase correction.
I turn on the option HyperComplex in the followig FTs just because I plan to eventually revise the phase correction along f-3. Otherwise I never use this option.
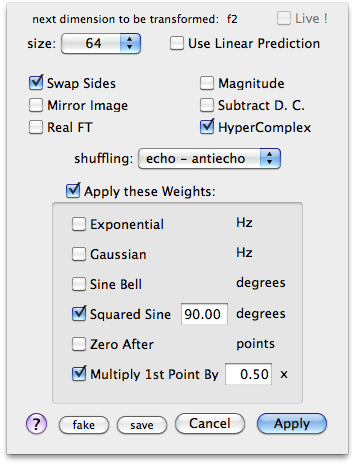
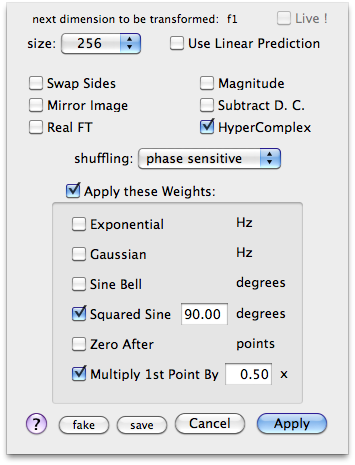
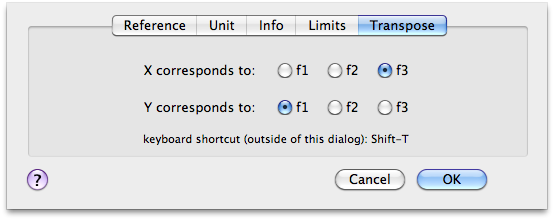
After the third FT we are in frequency domain. I transpose the cube to observe the f1-f3 planes. The command Format > Axes & Scales performs the transposition. I use the keys r and t to move through the planes. Now I can correct the phase along f-1 and f-3. When all is OK, I transpose again to correct the phase along f-2.
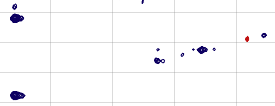
Actually this spectrum needs no phase correction at all along neither f-1 nor f-2. It is impossible to correct the phase of all the peaks without using the first-order correction. I can tolerate that the two peaks at the far left remain slightly out-of-phase. All the other peaks are phase-corrected.
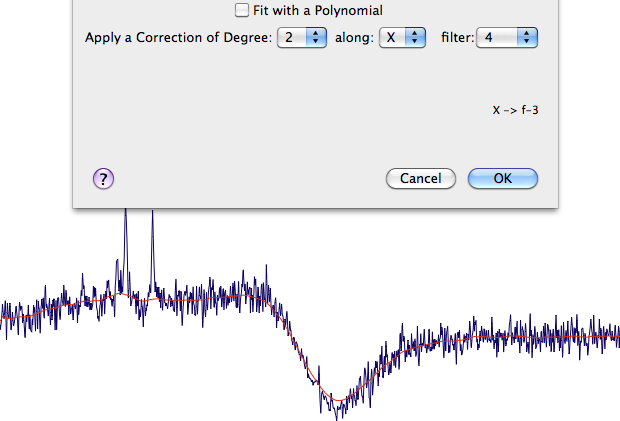
Now it would be great to remove the residual tails of the water signal that have resisted to the suppression algorithm. We can use the smoother algorithm in frequency domain, which can remove any broad signal. The problem is that, if this tool is not used with care, it can smooth the good peaks too. I extract a plane (containing several peaks), then an f-3 row, then I open the dialog Process > Automatic Baseline. Now I can verify which correction respects the good peaks while removing the unwanted ones. First I find the optimal Degree, then the optimal filter. After clicking OK, to store these values, I close all the extracts and apply the same command to the whole matrix.
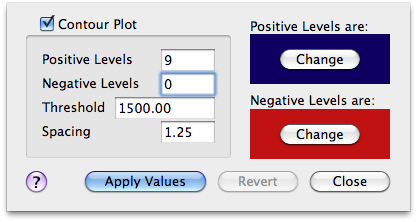
All the peaks of interest have a positive sign. If I hide the negative contour levels I remove half of the noise!
To increase the resolution you can also try to substitute Linear Prediction for Zero-Filling, for the indirect dimensions only. Avoid the “use Linear Prediciton” option of the FT dialog. It is very fast, but not very stable. Use, instead, the command Process > Linear Prediction and choose “Singular Value Decomposition” with, for example, “Number of Signals” = 4. SVD-LP is much slower but more stable, numerically speaking.
The file containing this spectrum is quite bulky (26 MB). You can download it, if you like, and repeat the tutorial by yourself.
