How to Fit a Portion of a Spectrum:

Process the experimental data: correct the phase and the baseline, calibrate the frequency axis and increase the resolution, if necessary to resolve the small couplings.
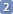
Create a simulated spectrum and perform a preliminary manual fitting.
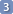
Now it's time to take some strategical decisions. At this stage you can start deciding which parameters are already correct (because you have measured them from a clear first order multiplet or because they are in accordance with the literature or your experience) and which remain unknown (they require optimization by the computer). Also choose the battlefield: which cluster of peaks to fit first. We suggest going from the simplest to the most complex one. Select and expand the first (or only) cluster of peaks.
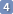
The sidebar contains a list of parameters. Put a check mark on the chemical shifts and the couplings of the nuclei involved. Don't check the parameters that you want to remain constant. As a general rule, it's more effective to optimize n parameters together than to optimize them one by one. Leave out the parameters that have nothing in common with the cluster under study.
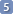
Check any of the population parameters if you also want to recalculate the populations, but if you have already done it (during the manual fitting) it's better to keep them constant. During the initial stage of the optimization, it is not necessary to simulate the authentic line width. Set a value approximatively similar (or slightly smaller).
Choose Simulate > Fit to Overlay. The visible part of the spectrum will be fitted. This command not only minimizes the difference, it also cuts away the regions of the simulated spectrum that are not visible. If you have fragmented the scale with the cutter, only the rightmost fragment will be considered. At the end of the calculation, before moving to another region of the spectrum, click the button refresh to restore the missing parts.
If the first attempt fails, increase the parameter pull and try again.
If the two corresponding regions are similar enough, keep constant the parameters adjusted so far and fit the other regions. Save the document often, so you can restore it if something goes wrong.
When all the multiplets are roughly similar, you can set pull = 0 and try simulating the line widths. This step is seldom necessary, but can improve the overall accuracy. If the shape of the experimental peaks is not purely lorentzian (because you have weighted the FID), you should also change the parameter %Lor (or let iNMR optimize it).

In the end, when you have found fair values for all the parameters, and if you have time to spare, you may save and experiment new strategies.